The 2015 qBio Bootcamp is over, and it was a lot of fun! Take a look below for some of the things we did.
Pictures from the camp
Teaching Assistants
Polly Parks, Department of Biology, SDSU
Steven Brown, Department of Physics and Molecular Biology, UCSD
JT Sauls, Bioinformatics and Systems Biology, UCSD
Participants
Suyeon Ju, ju.suyeon71@gmail.com, Scripps Ranch HS
Samantha Mendoza, smendoza2525@gmail.com, Mission Vista HS
Matthew Humkey, mhumkey33@gmail.com, Miramar College
Arsen Golemi, Miracosta College
Yamen Mubarka, yamen_mubarka@yahoo.com, UCSD
Bianca Luansing, bluansing@gmail.com, SD HS of International Studies
Alexa Nguyen, a_nguyen314@yahoo.com, Univ. of Arizona
Geoffrey Lu, geoffreylu@outlook.com, Rancho Bernardo HS
Byungjin Lee, wowbj08@naver.com, Chungnam National University
Nancy Meier, nancy.e.meier@gmail.com, UCSC
Guest Lecturers
Dr. Moselio Schaechter, Professor Emeritus of Biology, SDSU and UCSD
Dr. Stanley Maloy, Professor of Biology and Dean, College of Sciences, SDSU
Dr. Forest Rohwer, Professor of Biology, SDSU
Dr. Scott Rifkan, Professor of Biology, UCSD
Dr. Jing Xu, Professor of Biophysics, UC Merced
Dr. Michael Erikstad, Posdoctoral Scholar in Physics, UCSD
Dr. Mya Warren, Postdoctoral Scholar in Physics, UCSD
Overview and experiment
The bootcamp runs two weeks, from July 6th to July 17th. The first week (July 6th-10th) will be at Dr. Segall’s lab at SDSU. The second week (July 13th-17th) will be at Dr. Jun’s lab at UCSD. The camp runs all week M-F, from 9AM to 5PM, though we mostly likely will go late and perhaps meet on the intervening weekend, enthusiasm permitting.
General daily schedule:
9am: Introduction to and discussion of experiments
10am: Occasional guest speaker
12pm: Lunch
12:30pm+: Experimentation, data analysis, and write-up
4pm: Occasional guest speaker
Week 1
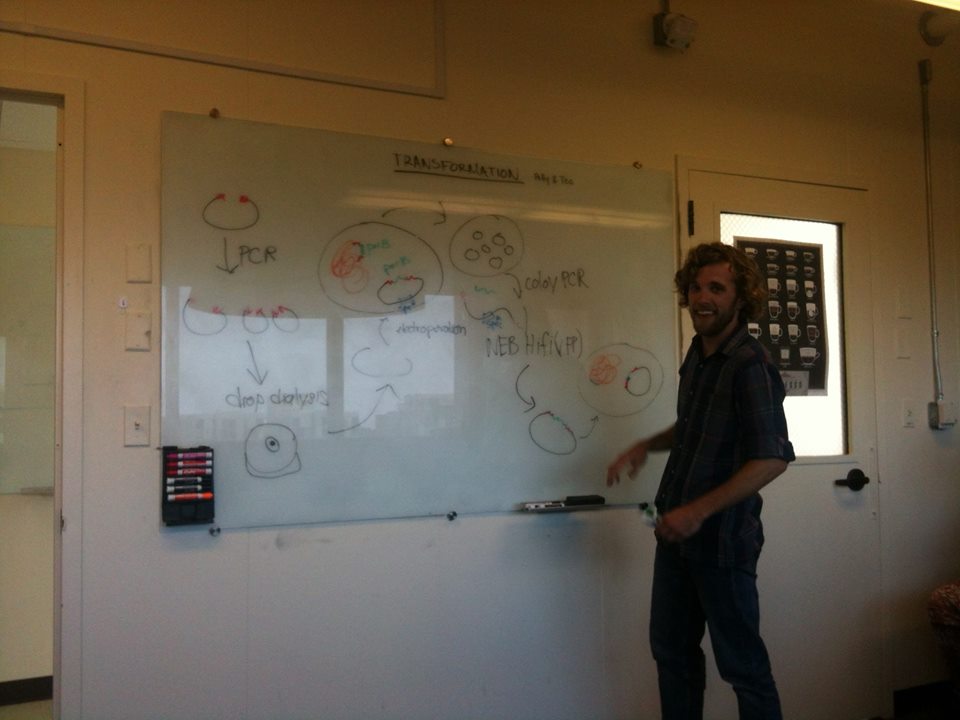
Week 1 will focus on genetics and construction of bacterial strains. We will design and build plasmids, transfer this genetic material/information into cells, and select for desired clones. Through these methods we will learn about the “axes and picks” of microbial genetics used to answer scientific questions. Technically, this involves:
Isolating microbial clones
Plasmid design and construction
Making DNA by PCR
Transforming cells by electroporation
Constructing strains by phage transduction
Genome modification using λ Red
Selection of bacterial mutants (Luria-Delbrück experiment)
Week 2
In week 2 we will grow and quantitatively study the bacterial strains we made in week 1. We will rely heavily on microscopy to understand their physiology–that is–their growth and biological processes. We will learn approaches to understanding quantitative data, what we can do with it, and how to effectively present our findings.
Growth curves via batch and continuous culture
Casting & assembling microfluidic devices
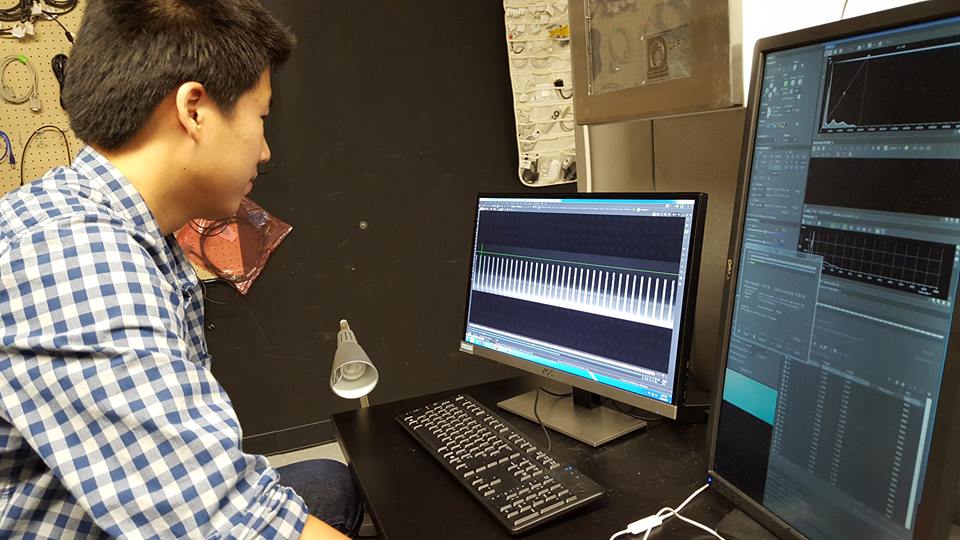
Capturing movies of single-cell growth
Fluorescent imaging with fusion proteins
Modeling mutation rates and the Luria-Delbrück experiment using python
Analyzing & interpreting image data sets
Summarizing & presenting information
Meals
Lunch, refreshments, and snacks will be provided. If we work late, we’ll get pizza. Try to enjoy a hearty breakfast before lab.
Computers
Some desktop computers will be provided but students are encouraged to bring their own laptops if possible.
Lab books, notes, protocols, and literature
Reading material, protocols, and notebooks will be provided. During the bootcamp, no prior reading is required though some material may be distributed the weekend before the camp for those interested.